A new GRIP on post-translational modifications
A highlight of Karaj*, Sindi*, Siriwongsup*, and Lee* et al. (2026) by Caroline Holley (*equal first authors).
NB: At the time of this highlight, the preprint is missing a materials and methods section, possibly due to intellectual property concerns over GRIP synthesis.
LINK TO ARTICLE: https://www.biorxiv.org/content/10.64898/2026.02.20.706349v1.full.pdf
BIORXIV DOI: 10.64898/2026.02.20.706349
ORIGINAL UPLOAD DATE: February 20th, 2026
Enzymes that modify proteins are rarely limited to a single substrate. Targeting these “writers/erasers” of post-translational modifications (PTMs) effectively rewires biological responses, but enzyme inhibition impacts the entire substrate pool and has other undesirable outcomes. A high-concern example in medicine is Janus Kinase (JAK) inhibitor discontinuation syndrome, where accumulation of phosphorylated but impotent JAK triggers lethal hyperactivity following inhibitor withdrawal (Andraos et al., 2012). This molecular whiplash limits the safety of JAK-inhibitor deployment against cancer and graft-versus-host disease. Tuning the writer/eraser interactome to target specific PTMs emerges as a safer strategy, or at least a necessary complement to existing therapies. Proximity-inducing chimeras that unite PTM writers/erasers with specific proteins of interest (POIs) already exist; PROteolysis TArgeting Chimeras (PROTACs) link POIs to ubiquinylation machinery (Békés et al., 2022), and similar strategies also target POIs to phosphatases (Fernandes et al., 2020). Proximity-inducing chimeras require an enzyme-binding handle (based on non-inhibitory and allosteric molecules), followed by a short linker and a POI-specific recognition domain. Compounds fitting these handle criteria are extremely rare, bottlenecking the application potential of proximity-inducing chimeras. In a recent BioRxiv preprint, Karaj, Sindi, Siriwongsup, and Lee et al. (2026) overcome these limitations by introducing first-in-class GRoup-transfer chimeras for Inducing Proximity (GRIPs). This work positions GRIPs as scalable tools for highly specific substrate targeting and extensive application across multiple domains of cell biology.
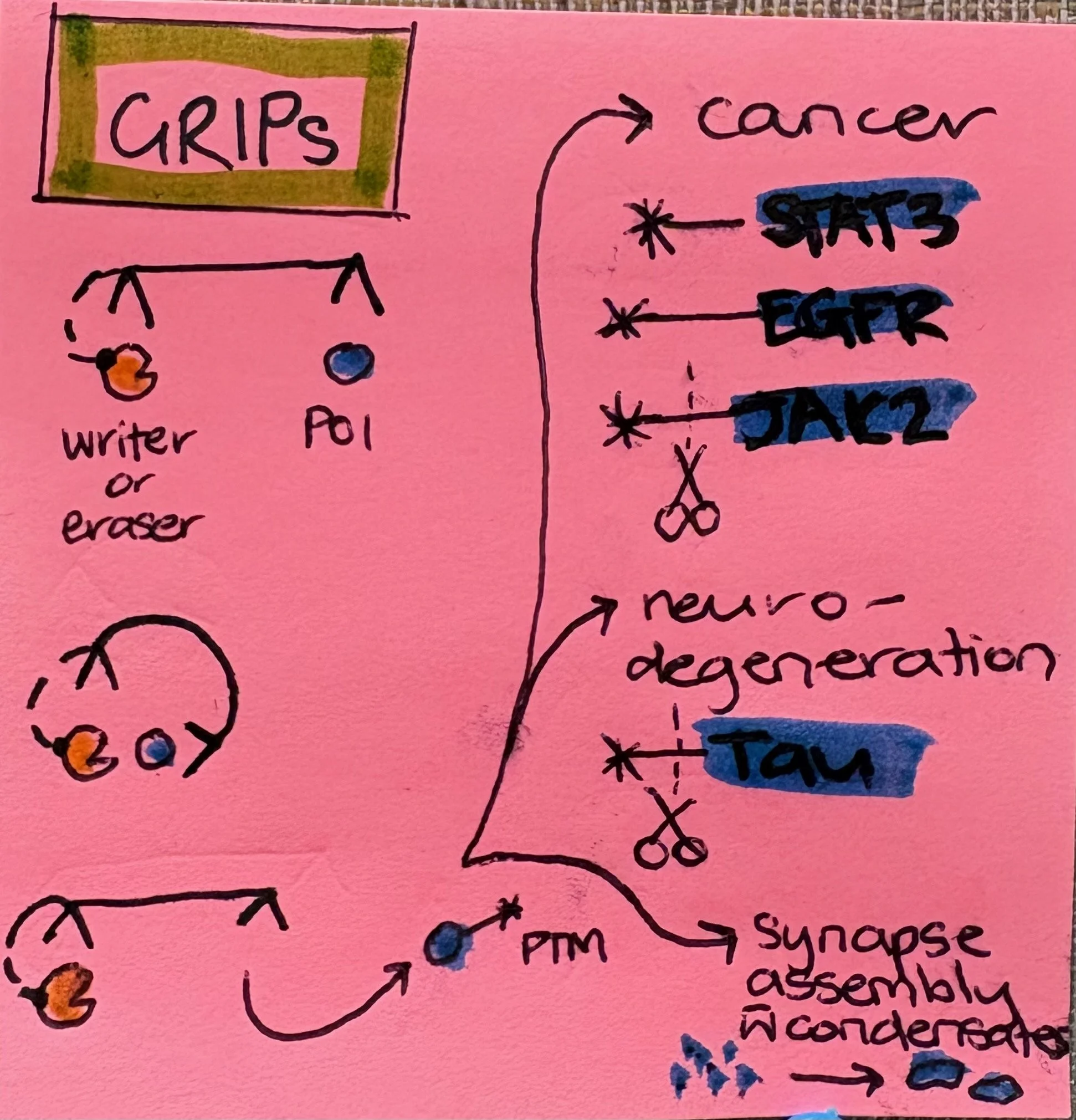
A post-it graphical abstract I drew in the time it took to finish one coffee.
The authors began by expanding writer/eraser handles beyond uncommon covalent and non-inhibitory molecules to more common non-covalent inhibitors. Writers/erasers with a proximal free cysteine near the inhibitor binding site were mined from the Protein Data Bank. This cysteine provides an anchor for a covalent group-transfer reaction, transforming what was initially a transient molecular handshake into solid handcuffs. Here, the first GRIP handle was modified from the non-covalent allosteric SHP099 inhibitor of SHP2 (Src homology-2-containing protein tyrosine phosphatase 2). In overexpression systems, an inducible dimerisation system was integrated into the POI-binding arm. Highly-specific inhibitors were sourced for endogenously-expressed POIs. This SHP2-GRIP system was validated across multiple substrates with roles in oncogenesis and neurodegeneration (STAT3, Tau, EGFR, and JAK2) under each of their pro-phosphorylating conditions. In each case, specific POI dephosphorylation was achieved without directly impacting other SHP2 targets. JAK2-targeting ablated the activity-rebound associated with withdrawal syndrome, providing an immediate proof-of-concept for GRIP utilisation as a complementary therapeutic. Interestingly, phosphatase efficiency at tyrosine versus serine/threonine residues was linker-dependent, not handle-dependent; an incidental finding and perhaps an exploitable layer for further tuning of target residue specificity. These data mark the first generation and success of an engineered GRIP handle from a non-covalent inhibitor.
Beyond SHP2 and dephosphorylation, GRIPs also successfully directed glycosylation. The glycosidase O-GlcNAc transferase (OGT; the writer) and O-GlcNAcase (OGA; the eraser) were both targeted endogenously. The same strategy for SHP2 was mirrored; non-covalent inhibitors of OGT/A with binding sites adjacent to a cysteine residue were identified. Only overexpressed POIs (casein kinase 2a, STAT5) were validated against the OGT/A GRIPs, perhaps owing to a lack of appropriate binding molecules against endogenous POIs. Again, linker-dependent efficiency was observed, highlighting the importance of strategic design beyond GRIP handles for different cellular applications.
Further scalability to other writers/erasers required further re-engineering of the GRIP handle. The group-transfer arm (that covalently binds the cysteine) was modified for compatibility with covalent inhibitors and also for lysine anchor residues. Engineering GRIP handles to accommodate lysine group-transfer reactions expanded the writer/enzyme repertoire, as lysines are more abundant in the proteome than free cysteines. These combined approaches allowed targeting of AKT (protein kinase B) and AMPK (AMP-activated protein kinase) to the Liprin-a3 POI, which induced assembly of cytosolic phase-separated condensates. Liprin-a3 condensates are critical signalling platforms for neuronal synapse assembly (Marcó de la Cruz et al., 2024), hinting at the potential for GRIPs to specifically rewire neuron circuitry. While molecular tethers already exist for inducible organelle-organelle contacts (Eisenberg-Bord et al., 2016), most require genetic encoding of an inducible dimeriser or a non-permeable protein scaffold. While this GRIP system still required overexpression of a recruitable Liprin-a3 fusion protein, all that remains for a fully-endogenous strategy is identification or synthesis of a specific Liprin-a3-binding molecule. These data are the first example of PTM writer/eraser proximity-inducing chimeras manipulating condensate assembly.
In a final example of GRIP utility, the authors answered the question: what if the writer and POI are the same protein? Epidermal Growth Factor Receptor (EGFR) can auto-phosphorylate in the absence of EGF (Schlessinger, 2004) and drive anti-tumourigenic programs. Here, the authors used the same EGFR inhibitor-based interaction domain mirrored on the GRIP handle and the POI-binding arm, successfully inducing dimerisation and autophosphorylation. As cancer-associated KRAS (Kirsten rat sarcoma virus oncogene homologue) mutant-bearing cells are pre-primed and already sensitive to EGFR manipulation, this EGFR GRIP was tolerated in wild-type cells but uniquely lethal to cells bearing KRAS mutations. This fully-endogenous approach further highlights the utility of GRIPs across cell biology and, if equally effective in vivo, medical application.
Karaj, Sindi, Siriwongsup, and Lee et al. (2026) presented first-in-class GRIPs for tuning the PTM-ome beyond the capabilities of simple inhibition. They first overcame the major limitation of generating diverse proximity-inducing chimeras by designing molecular handles compatible with covalent and non-covalent inhibitors. These GRIPs were immediately tested against therapeutically relevant targets (e.g. STAT3, JAK2) of phosphorylation and glycosylation. While not directly addressed or assayed, it appears all GRIPs were cell-permeable and most were efficacious at nanomolar concentrations. This system still requires having pairs (writer/eraser plus POI) of highly-specific inhibitors or binding molecules, which did restrict some experiments to overexpressed fusion protein POIs. The next step is perhaps generating an AI-assisted database of molecule pairs for customisable GRIP handle synthesis. Certainly there are still many targets to explore with current resources; many crystallisation studies employ inhibitors to achieve a high-resolution structure, and so the binding sites and inhibitor affinities are well-characterised. It will be interesting to next probe the in vivo efficacy of GRIPs, and whether GRIPs can be multiplexed with existing targeting strategies to execute cell-specific PTM manipulation.
REFERENCES
Andraos, R., Qian, Z., Bonenfant, D., Rubert, J., Vangrevelinghe, E., Scheufler, C., Marque, F., Régnier, C. H., De Pover, A., Ryckelynck, H., Bhagwat, N., Koppikar, P., Goel, A., Wyder, L., Tavares, G., Baffert, F., Pissot-Soldermann, C., Manley, P. W., Gaul, C., … Radimerski, T. (2012). Modulation of activation-loop phosphorylation by JAK inhibitors is binding mode dependent. Cancer Discovery, 2(6), 512–523. https://doi.org/10.1158/2159-8290.CD-11-0324
Békés, M., Langley, D. R., & Crews, C. M. (2022). PROTAC targeted protein degraders: the past is prologue. Nature Reviews Drug Discovery 2022 21:3, 21(3), 181–200. https://doi.org/10.1038/s41573-021-00371-6
Eisenberg-Bord, M., Shai, N., Schuldiner, M., & Bohnert, M. (2016). A Tether Is a Tether Is a Tether: Tethering at Membrane Contact Sites. Developmental Cell, 39(4), 395–409. https://doi.org/10.1016/J.DEVCEL.2016.10.022/ASSET/C97F4D08-FA51-4A54-A214-A9305E8874F8/MAIN.ASSETS/GR4.JPG
Fernandes, R. A., Su, L., Nishiga, Y., Ren, J., Bhuiyan, A. M., Cheng, N., Kuo, C. J., Picton, L. K., Ohtsuki, S., Majzner, R. G., Rietberg, S. P., Mackall, C. L., Yin, Q., Ali, L. R., Yang, X., Savvides, C. S., Sage, J., Dougan, M., & Garcia, K. C. (2020). Immune receptor inhibition through enforced phosphatase recruitment. Nature, 586(7831), 779–784. https://doi.org/10.1038/S41586-020-2851-2
Marcó de la Cruz, B., Campos, J., Molinaro, A., Xie, X., Jin, G., Wei, Z., Acuna, C., & Sterky, F. H. (2024). Liprin-α proteins are master regulators of human presynapse assembly. Nature Neuroscience 2024 27:4, 27(4), 629–642. https://doi.org/10.1038/s41593-024-01592-9
Schlessinger, J. (2004). Common and distinct elements in cellular signaling via EGF and FGF receptor. Science, 306(5701), 1506–1507. https://doi.org/10.1126/SCIENCE.1105396/ASSET/6DFBABC1-0191-4413-A66B-0C1D61A77485/ASSETS/GRAPHIC/306_1506_F1.JPEG
AI USE DISCLOSURE
Generate AI was not used for synthesising text in this highlight. ChatGPT5.2 was prompted to perform a final grammar and spelling check (Prompt: check the attached text for spelling and grammar mistakes. Suggest corrections but do no not alter the text in any other way”).
