The P-Body Problem: a super NOVA reveals ALS-associated organelle dysfunction
A highlight of Krispin et al. (2026) by Caroline Holley
LINK TO ARTICLE: https://www.biorxiv.org/content/10.1101/2024.01.31.572110v3.full.pdf
BIORXIV DOI: 10.1101/2024.01.31.572110v3
ORIGINAL UPLOAD DATE: January 21st, 2024 (revised February 2nd, 2026)
Amyotrophic lateral sclerosis (ALS) is a devastating neurodegenerative disease with rapid onset, deterioration, and certain mortality. Patients experience progressive loss of motor control, culminating in failure of essential bodily functions (mobility, ability to swallow, and breathing). There is no cure and no treatment that slows progression. The precise cellular events preceding disease onset are also poorly understood, and only 10% of ALS cases can be traced to genetic risk factors (Nijs & Van Damme, 2024). In a recently-revised preprint, Krispin et al. (2026) develop a high-throughput learning model to analyse organelle perturbations across almost every cellular compartment under ALS-associated oxidative and genetically encoded stress. As a model for image analysis, this manuscript represents a major leap forward in the complex analysis of diverse imaging data sets that handles highly heterogeneous, close-to-primary cell data robustly, with the ability to accurately report interactions between networks of associated organelles and assist in dissecting causal hierarchy. For the ALS case, this manuscript applies this model to uncover the role of a newly characterised RNA-processing organelle, P-bodies, in disease onset and supports these conclusions with real patient-derived induced pluripotent stem cell-derived (iPSC) neurons and post-mortem tissue samples.
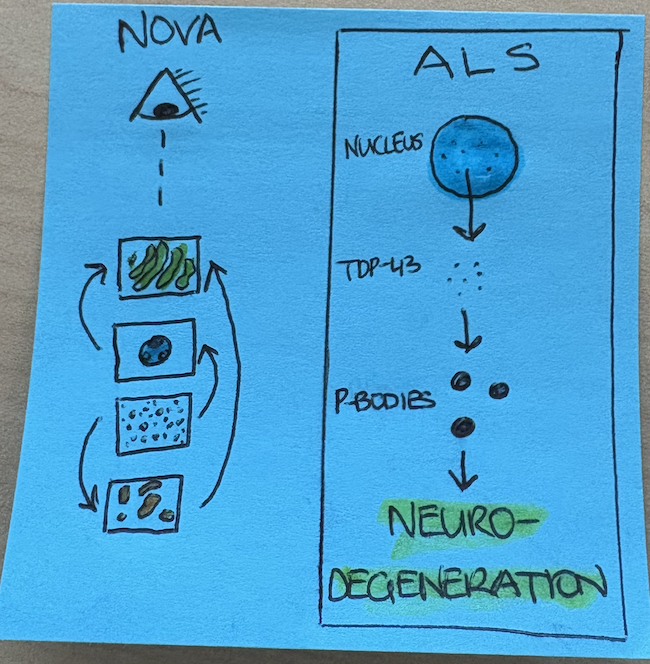
A post-it graphical abstract I drew in the time it took to finish one coffee.
Most people are familiar with transformer architecture in public-facing language learning models, the most notable example being ChatGPT. Transformer models are ideal for making sense of complex and distant relationships between variables, and here the authors have applied a vision transformer (ViT; which handles images rather than language) architecture to classifying the cellular topography of organelles in human iPSC-derived neurons. The model, called “NOVA”, was trained on an internally generated data set of neurons stained for about 26 different organelles and compartments under resting or chemically perturbed conditions. iPSC neurons are highly heterogeneous, presenting both a challenge in training and the eventual payoff of a robustly trained model compared to those trained on small or homogeneous datasets from routinely used immortalised cell lines. NOVA was trained to control for natural variation in organelle shape and localisation and was >99% accurate at organelle identification. This is particularly impressive given how many organelles exhibit similar and/or highly heterogeneous morphology (e.g., punctate condensates or vesicles). Interestingly, the model identified distinct differences between Mitotracker and mitochondrial outer membrane staining; however, this aligns with anecdotal evidence from myself and others that Mitotracker promiscuously stains other non-mitochondrial membranes. Treatment of neurons with sodium arsenite, a chemical agent that causes oxidative stress (one of the known hallmarks of ALS), produced several statistically significant organelle perturbations. Of these, stress granules and P-bodies emerged with the highest effect shift. P-bodies, or “processing bodies”, are mRNA-rich condensates in the cytoplasm that are enriched for several RNA-processing enzymes. In iPSC neurons engineered to carry genetic variants associated with ALS (10% of ALS cases), most assayed organelles were perturbed, revealing a global disruption of organelle homeostasis, but the precise organelle signatures depended on the nature of the genetic ALS predisposition. A few were more prone to stress granule disruption, while most other backgrounds recapitulated the P-body disruption observed earlier with chemically induced oxidative stress. There was also a strong lysosomal phenotype, supporting endolysosomal associations with the ALS genetic markers (Rosso et al., 2026). Here, NOVA proved useful in linking organelle–organelle relationships to identify true perturbations versus indirect effects due to an interacting organelle, uncovering very subtle changes that would be missed by eye or conventional image analysis.
Another hallmark of ALS, observed across cell models and post-mortem tissue, is TDP-43 mislocalisation from the nucleus to the cytoplasm (Trist et al., 2022). While some heritable ALS cases, including several assayed in this manuscript, trigger TDP-43 mislocalisation and aggregation, TDP-43 aggregation is also significantly upregulated in iPSC neurons cultivated from “spontaneous” patients without an explainable genetic predisposition (Rothstein et al., 2025). The resulting TDP-43 aggregates may disrupt autophagy (Dopler et al., 2025), and extracellular TDP-43 has inflammatory signalling consequences (Evangelista et al., 2025). It is also known that ALS patients have defective mRNA processing and that TDP-43 is an RNA-binding protein that co-regulates mRNA stability (Alessandrini et al., 2025), but the precise link between TDP-43 and disease onset is not clear. Indeed, several of the genetic markers assayed earlier in the manuscript triggered TDP-43 accumulation in the cytoplasm. Here, the authors forced TDP-43 cytoplasmic accumulation by removing its native nuclear localisation sequence (TDP-43ΔNLS). TDP-43ΔNLS iPSC neurons accumulated TDP-43 in the cytoplasm, and P-bodies and PML (promyelocytic leukemia; nuclear) bodies were significantly disrupted. PML bodies may regulate TDP-43 (Wagner et al., 2025), and these data have uncovered a previously unknown reverse relationship. Imaging of P-bodies and TDP-43 revealed co-localisation, supported by proximity labelling of TDP-43 and highly enriched P-body hits following enrichment of labelled interactors. Phenotypically, P-bodies in TDP-43ΔNLS appeared smaller, and FRAP experiments revealed that these condensates were compressed and less fluid compared to those in WT TDP-43-expressing U2OS cells. This was only observable in U2OS cells upon chemically induced oxidative stress, likely representing a quirk of the U2OS model rather than true physiology, as this was not required in iPSC neurons. As P-bodies are key players in mRNA processing, mRNA dynamic properties were examined in TDP-43ΔNLS neurons; 30% of P-body-associated mRNAs were destabilised, and 5% of global mRNAs. The authors do not identify what these other mRNAs are; however, it would be intriguing to follow up.
Now that NOVA has uncovered multiple organelle perturbations using different models of ALS, the authors next sought to confirm these data in primary ALS patient samples. TDP-43 and P-bodies in ALS patient-derived iPSC motor neurons were imaged and analysed using NOVA. Of the originating patients, half exhibited characteristic cytoplasmic TDP-43, and most patients had spontaneous ALS. Primary cells revealed additional nuclear and microtubule effects on top of the P-body and granule disruption observed in the engineered models, but the TDP-43 and P-body colocalisation was conserved when TDP-43 localisation was disrupted. Patient organelle effects were biased towards endolysosomal dysfunction rather than P-bodies compared to the lab-standard iPSC neurons in some genetic backgrounds, but P-body reduction was significant or nearly significant across other patient cohorts as long as they shared TDP-43 mislocalisation. In post-mortem brain samples, TDP-43 colocalised to varying degrees with P-bodies, and this variability is perhaps explained by sample preparation or storage conditions of post-mortem tissue not being as controlled as iPSCs. Mirroring the earlier models, patients with TDP-43-depleted nuclei had lower P-body mRNA levels, with statistical significance varying between donors.
In sum, the authors have developed a high-throughput intelligent image analysis model that handles complex organelle relationships to reveal cryptic features of cell biology in disease contexts. The NOVA model itself may also prove useful in unrelated studies of phase-separated condensates, as these can only really be studied by imaging and likely have labile interactions with other organelles and compartments. This manuscript represents a significant technical advancement in AI-driven image analysis and an equally impactful biological advance in understanding the cellular events that accompany ALS disease. While TDP-43 mislocalisation was not triggered in all the genetic backgrounds here, it is a conserved feature of many unexplained spontaneous ALS cases. These data and this tool may go a long way in unravelling the cryptic cell biology of this devastating disease. These data prompt further questions: how exactly are dysfunctional P-bodies derailing healthy neuronal function to drive ALS? And can we stabilise P-bodies to rescue neuronal dysfunction and cell death in ALS?
AI USE DISCLOSURE
I wrote this highlight myself. In addition to manually searching literature on PubMed and Google, I utilised OpenSci Ai2 (prompts: “Find papers on ALS, P-bodies, and TDP-43” and “There are genetic and spontaneous ALS cases. In how many spontaneous ALS cases is TDP-43 mislocalised or aggregating in the cytoplasm?”). I used ChatGPT 5.2 to assist with proof-reading (prompt: “suggest grammar edits”)
REFERENCES
Rothstein, J. D. et al. Sporadic ALS induced pluripotent stem cell derived neurons reveal hallmarks of TDP-43 loss of function. Nature Communications 2025 16:1 16, 7092- (2025).
Alessandrini, F. et al. TDP-43 Dysfunction Compromises UPF1-Dependent mRNA Metabolism in ALS. Neuron S0896-6273(25)00848–7 (2025) doi:10.1016/J.NEURON.2025.11.001.
Trist, B. G. et al. Co-deposition of SOD1, TDP-43 and p62 proteinopathies in ALS: evidence for multifaceted pathways underlying neurodegeneration. Acta Neuropathol. Commun. 10, 122- (2022).
Evangelista, B. A. et al. ALS-associated TDP-43 aggregates drive innate and adaptive immune cell activation. iScience 28, (2025).
Wagner, K. et al. Induced proximity to PML protects TDP-43 from aggregation via SUMO–ubiquitin networks. Nature Chemical Biology 2025 21:9 21, 1408–1419 (2025).
Rosso, F. et al. Non-cell autonomous autophagy in amyotrophic lateral sclerosis: A new promising target? Neurobiol. Dis. 218, 107203 (2026).
Nijs, M. & Van Damme, P. The genetics of amyotrophic lateral sclerosis. Curr. Opin. Neurol. 37, 560–569 (2024).
Krispin, S. et al. Organellomics: AI-driven deep organellar phenotyping reveals novel ALS mechanisms in human neurons. bioRxiv 2024.01.31.572110 (2026) doi:10.1101/2024.01.31.572110.
Dopler, M. B. et al. A cellular model of TDP-43 induces phosphorylated TDP-43 aggregation with distinct changes in solubility and autophagy dysregulation. FEBS J. 292, 4870–4897 (2025).
